|
Sunday, January 24, 2010
Frequency of lactose malabsorption among healthy southern and northern Indian populations by genetic analysis and lactose hydrogen breath and tolerance tests:
Volunteers from southern and northern India were comparable in age and sex. The LTT result was abnormal in 88.2% of southern Indians and in 66.2% of northern Indians...The lactose HBT result was abnormal in 78.9% of southern Indians and in 57.1% of northern Indians...The CC genotype was present in 86.8% and 67.5%...the CT genotype was present in 13.2% and 26.0%...and the TT genotype was present in 0% and 6.5%..of southern and northern Indians, respectively. The frequency of symptoms after the lactose load...and peak concentrations of breath hydrogen...both of which might indicate the degree of lactase deficiency, were higher in southern than in northern Indians. 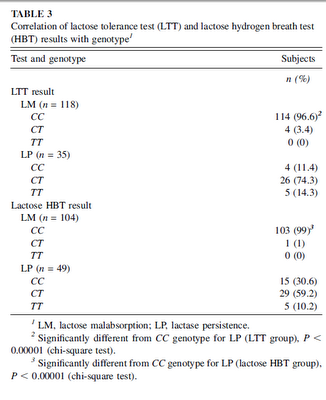 The north Indian samples were from Lucknow on the mid-Gangetic plain, and the south Indian samples from Bangalore. The genetic variant conferring lactase persistence is the Central Asian one, T-1390. You can see the distribution of the genotypes by phenotype in the table to the left. These authors assume that the T allele was brought by the Indo-Aryans; this seems plausible seeing its clinal variation, as well the fact that this variant seems to be common in European and Central Asian populations. The frequency of the T allele in the Lucknow sample was 39%, and 13% in the Bangalore sample. Here are a selection of frequencies for the T allele in other populations: The north Indian samples were from Lucknow on the mid-Gangetic plain, and the south Indian samples from Bangalore. The genetic variant conferring lactase persistence is the Central Asian one, T-1390. You can see the distribution of the genotypes by phenotype in the table to the left. These authors assume that the T allele was brought by the Indo-Aryans; this seems plausible seeing its clinal variation, as well the fact that this variant seems to be common in European and Central Asian populations. The frequency of the T allele in the Lucknow sample was 39%, and 13% in the Bangalore sample. Here are a selection of frequencies for the T allele in other populations:17% - Saami You can see more here. This looks like a case of local adaptation. Labels: India, Population genetics
Friday, October 30, 2009
Something that has been nagging me about the recent paper by Reich et al. which models Indian populations as a hybridization event between two ancestral groups, "Ancestral South Indians" (ASI) and "Ancestral North Indians" (ANI). As a reminder, the ANI seem to have been rather like Europeans in their allele frequencies, or at least far closer to Europeans than they were to the ASI (it seems that they compared ANI with Western Europeans). This is interesting. They found in the populations surveyed that the low bound for ANI was 40%, the high ~80% (in the supplements they included some Pathans and Sindhis from the HGDP, and that's where that number comes from). The ~40% low bound for ANI rather surprised me. The populations which they sampled included South Indian tribal groups. In other words, these were the groups arguably least affected by what we term Hinduism and Indian culture (their status as "tribals" as opposed to lower caste or outcaste was generally a function of the fact that they rejected integration and assimilation into mainstream Indian culture and isolated themselves both geographically and in terms of their customs). Just seems weird that these groups would be so ANI.
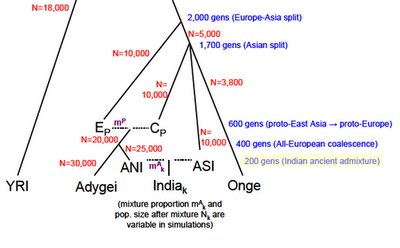 For a few weeks now Greg Cochran has been asking if I saw something in the paper above about when the admixture between ANI and ASI occurred, or at least if there was a hint about when the authors think it occurred. I said no, there are only hints. I was wrong, I skimmed over the supplement too quickly, they assume 200 generations ago as a parameter in a model they use for simulations. Bingo. Just click the image to the left, and look at the lower right. 200 generations = 5,000 years ago, assuming 25 years for generation time. Let's assume that a South Indian tribal group is a small deme of ASI surrounded by a very large (infinite) deme of ANI for 200 generations. If I assume a constant outmarriage rate of 0.25% per generation (1 out of 400) then at the present time you'd have the tribal group being ~40% ANI. For a few weeks now Greg Cochran has been asking if I saw something in the paper above about when the admixture between ANI and ASI occurred, or at least if there was a hint about when the authors think it occurred. I said no, there are only hints. I was wrong, I skimmed over the supplement too quickly, they assume 200 generations ago as a parameter in a model they use for simulations. Bingo. Just click the image to the left, and look at the lower right. 200 generations = 5,000 years ago, assuming 25 years for generation time. Let's assume that a South Indian tribal group is a small deme of ASI surrounded by a very large (infinite) deme of ANI for 200 generations. If I assume a constant outmarriage rate of 0.25% per generation (1 out of 400) then at the present time you'd have the tribal group being ~40% ANI.OK, what about my idea which I presented to John Hawks that Indians "don't really look" like a hybridization between Northern Europeans and the ASI, ASI assumed to be similar to the Andaman Islanders (who I do not believe were necessarily "Negritos," insofar as I suspect their small stature is due to contact with Europeans and Indians, as those who have avoided such contact are seen to be of normal or even above average size for South Asians). Specifically the frequency of light eyes and hair is just way too low among groups which are on the 70-80% ANI range such as Punjabis and Kashmiris, though these groups do tend have more Caucasoid features and lighter (olive) skin. On the other hand, here is something which jumped out at me about the Reich et al. paper: they added two Pakistani populations who fit well in the ANI-ASI cline which most of the Indian groups mapped onto (some groups with "Eastern" origin in both Pakistan and India were discarded from the analysis), and their ANI frequency proportions seemed familiar to me. There are three ANI estimates for both groups: Sindhi - 78%, 70.7%, 73.7% (78%) Pathan - 81%, 74.2%, 76.9% (81%) In the parenthesis is the frequency for the derived (European-like) variant of SLC24A5. The data sets were the same, from the HGDP, though the ancestry estimates used only 10 and 15 of the approximately 50 of each group respectively. There's a suspicious correspondence here. The lowest frequency of the derived variant of SLC24A5 I've seen for a South Asian population is ~30% for Sri Lankan Tamils, with ~50% for Sri Lankan Sinhalese. Remember that a reasonable low bound for ANI for South Asian groups is on the order of 40%. But what about my contention that other European-like pigmentation alleles don't fit because the phenotype isn't what you'd expect. You can look at a blue vs. brown eye variant of OCA2 in the HGDP. Another eye color variant, HERC2. And here is a variant of TYR which causes light skin. The interesting point would be to look at the Indian samples, but I don't have really good proxies for that (in one paper which surveyed Indian Americans various language groups ranged from 70-100% in derived SLC24A5 frequency, but it is very difficult to imagine that these correspond well to many groups in the Reich paper. Specifically, it's biased toward higher status/caste groups). I might have spoken too soon, though it still seems to me that something is off. Perhaps Europeans changed after ANI left. Or perhaps ANI changed when it arrived in India. One recent data point which I find curious is that a paper just came out which suggests that populations of the Andronovo culture in Trans-Siberia, which is assumed to be the precursor to the Indo-Iranians, seem to resemble modern day Russians in pigment phenotype. At least judging from the genes extracted and sequenced. More later when my thoughts become more settled. Labels: India, Population genetics |



